CPAP/BIPAP
Have you been diagnosed with obstructive sleep apnea? We can help!
Sleep better and breathe easier with Owens.
Learn about the therapies and services available through Owens.
Patient care is our top priority. Trust Owens to provide support and guidance every step of the way. State-of-the-art medical equipment, compassionate care, and 24/7 support are just a few of the reasons why patients and providers pick Owens.
Make an Appointment
Click here to reorder PAP supplies or Talk to specialist.
Check the status of your order online. Fast, easy, and secure
Shop Owens Direct online for the best prices and superior customer service.
Need to pay your bill? Click here to visit our secure payment site.

Owens professionals work closely with referring physicians and hospital staff to ensure seamless continuity of care from the hospital or doctor’s office into your home.
Learn MoreStop waiting on hold. Make the switch to Owens.
Confidence is knowing you will always be able to connect with our team of local medical equipment specialists during business hours, whether it’s about your patient’s unique healthcare needs or the specialty products and services we offer. We can answer questions about potential drug interactions, compliance requirements, and schedule follow-up services that take place in the patient’s home. We accept most insurance plans.
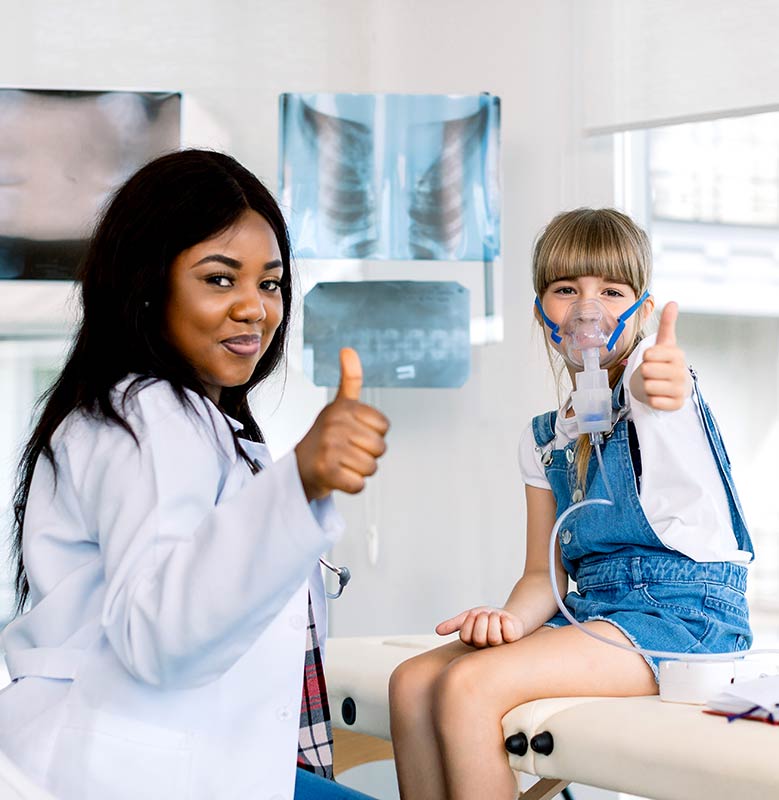

We’re committed to providing state-of-the-art equipment and solutions that assist you in treating patients with chronic respiratory needs.
Providing superior value and unmatched service throughout the North State.


Owens partners with leading health plans to provide cutting-edge technology solutions. With decades of contract experience, and full accreditation by the Joint Commission on Accreditation of Healthcare Organizations (JCAHO) since 2003, we’re providing superior value and unmatched service throughout the North State.
Owens has been locally owned and operated since 1957, when our first retail pharmacy was established in downtown Redding. Although we’ve continued to grow in the decades since, our dedication to the local community and commitment to providing the best-possible customer service has never waivered.
Learn about Owens

Enjoy fast, friendly service from a dedicated team of respiratory therapists and sleep specialists. Integrity, quality, and service are at the heart of everything we do. Owens has been locally owned and operated in Northern California since 1957.